(See Google Scholar for the Up-to-Date List)
Publications
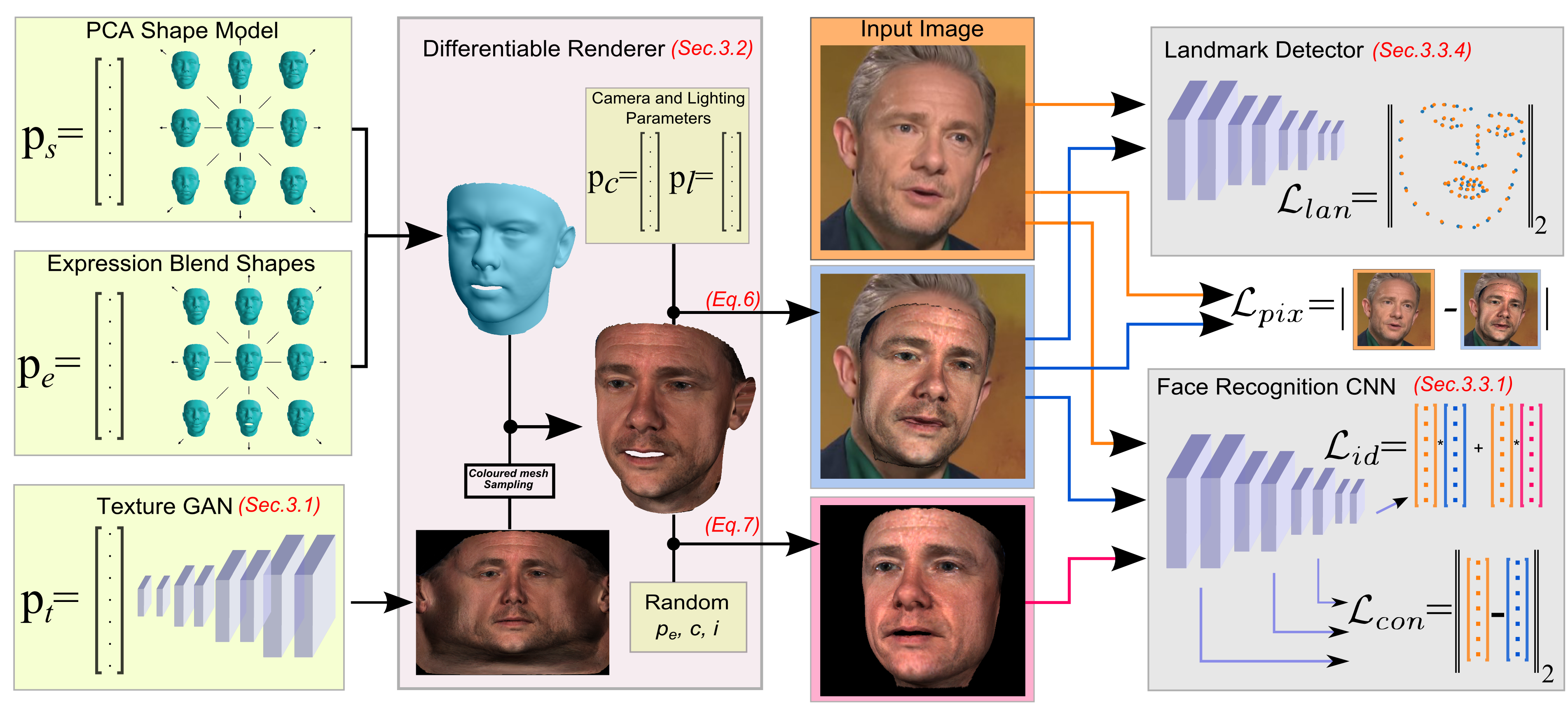
B. Gecer, S. Ploumpis, I. Kotsia, S. Zafeiriou, “GANFIT: Generative Adversarial Network Fitting for High Fidelity 3D Face Reconstruction” CVPR, 2019.
B. Gecer, B. Bhattarai, J. Kittler and TK. Kim, “Semi-supervised Adversarial Learning to Generate Photorealistic Face Images of New Identities from 3D Morphable Model” ECCV, 2018.
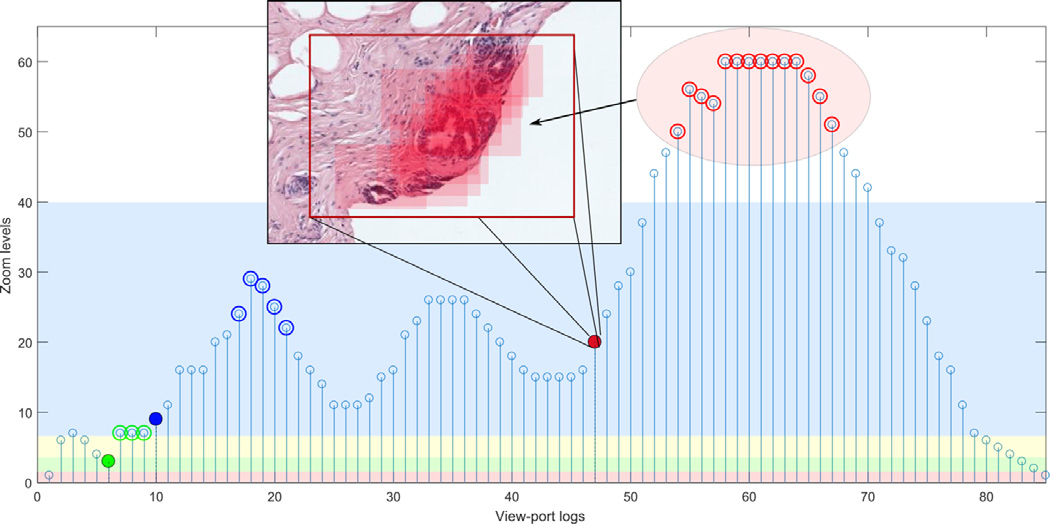
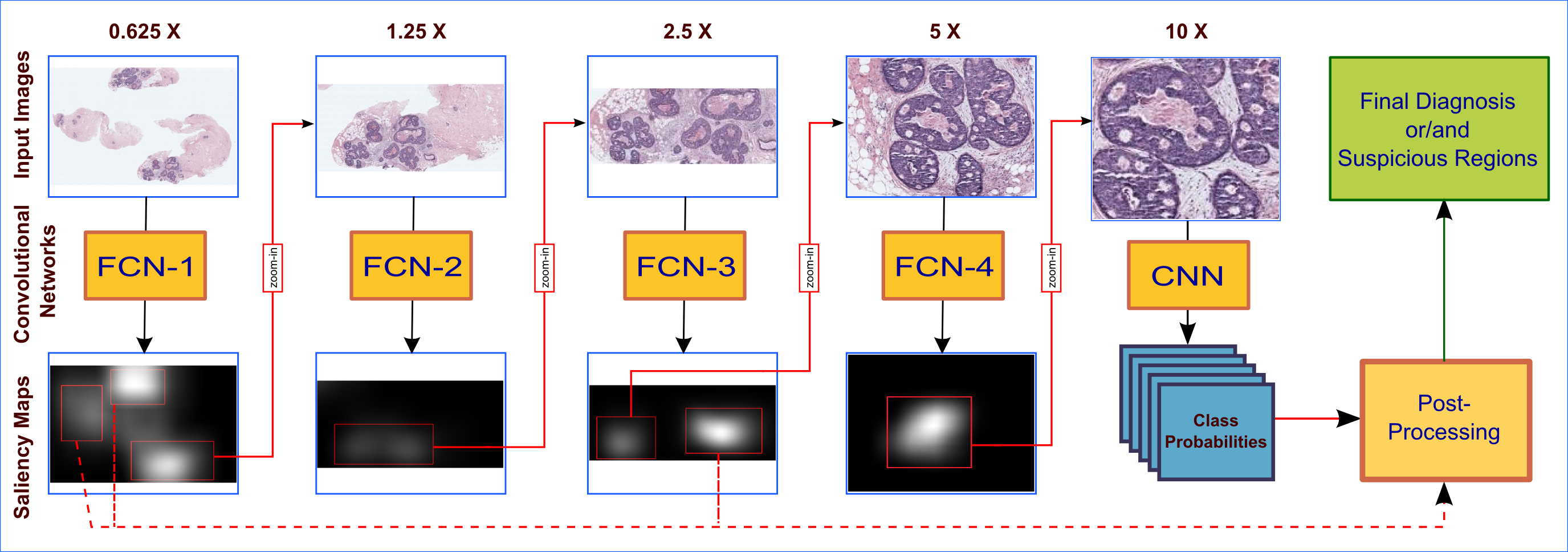
B. Gecer, S. Aksoy, E. Mercan L.G. Shapiro, D.L. Weaver and J.G. Elmore, “Detection and classification of cancer in whole slide breast histopathology images using deep convolutional networks” Pattern Recognition, 2018.
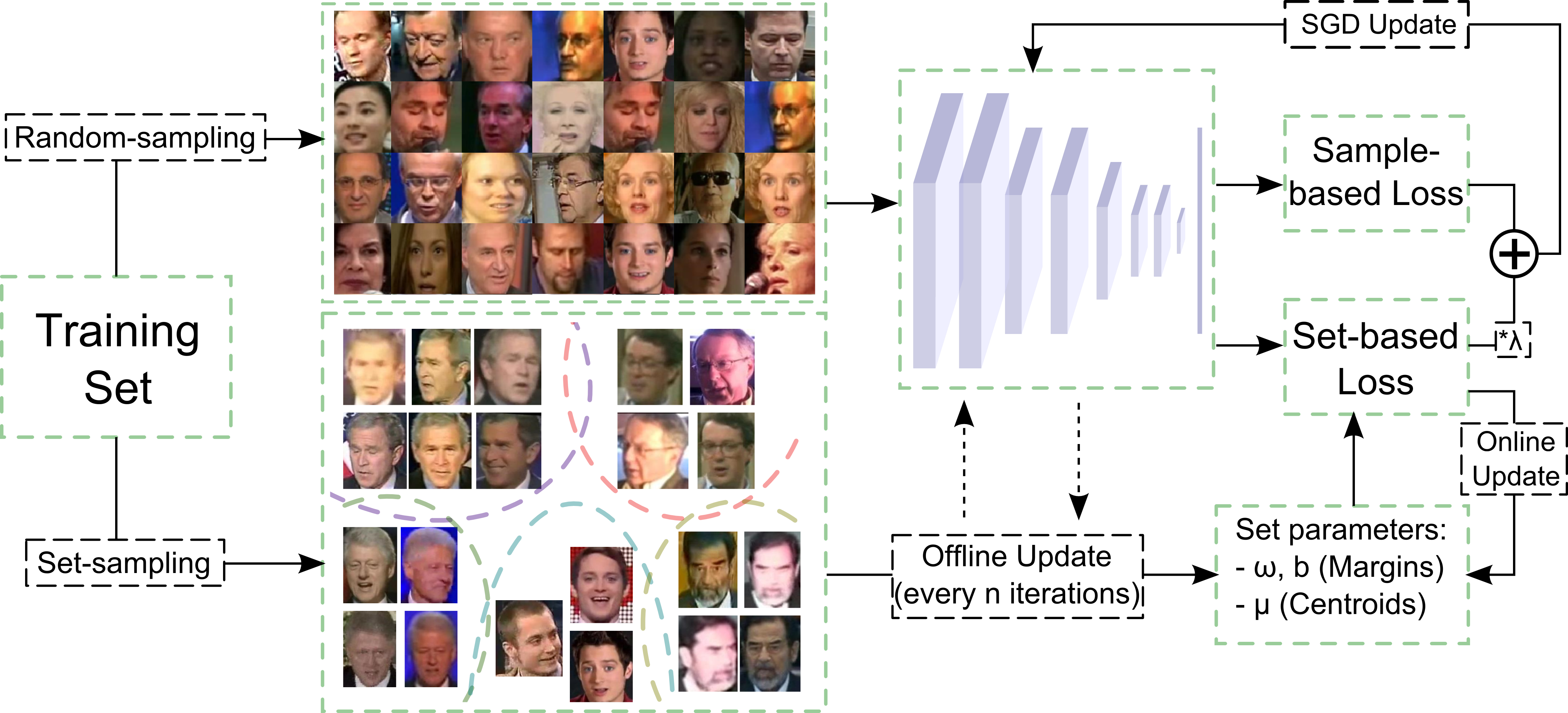
B. Gecer, V. Balntas, and TK. Kim, “Learning Deep Convolutional Embeddings for Face Representation Using Joint Sample- and Set-based Supervision” ICCVW, 2017
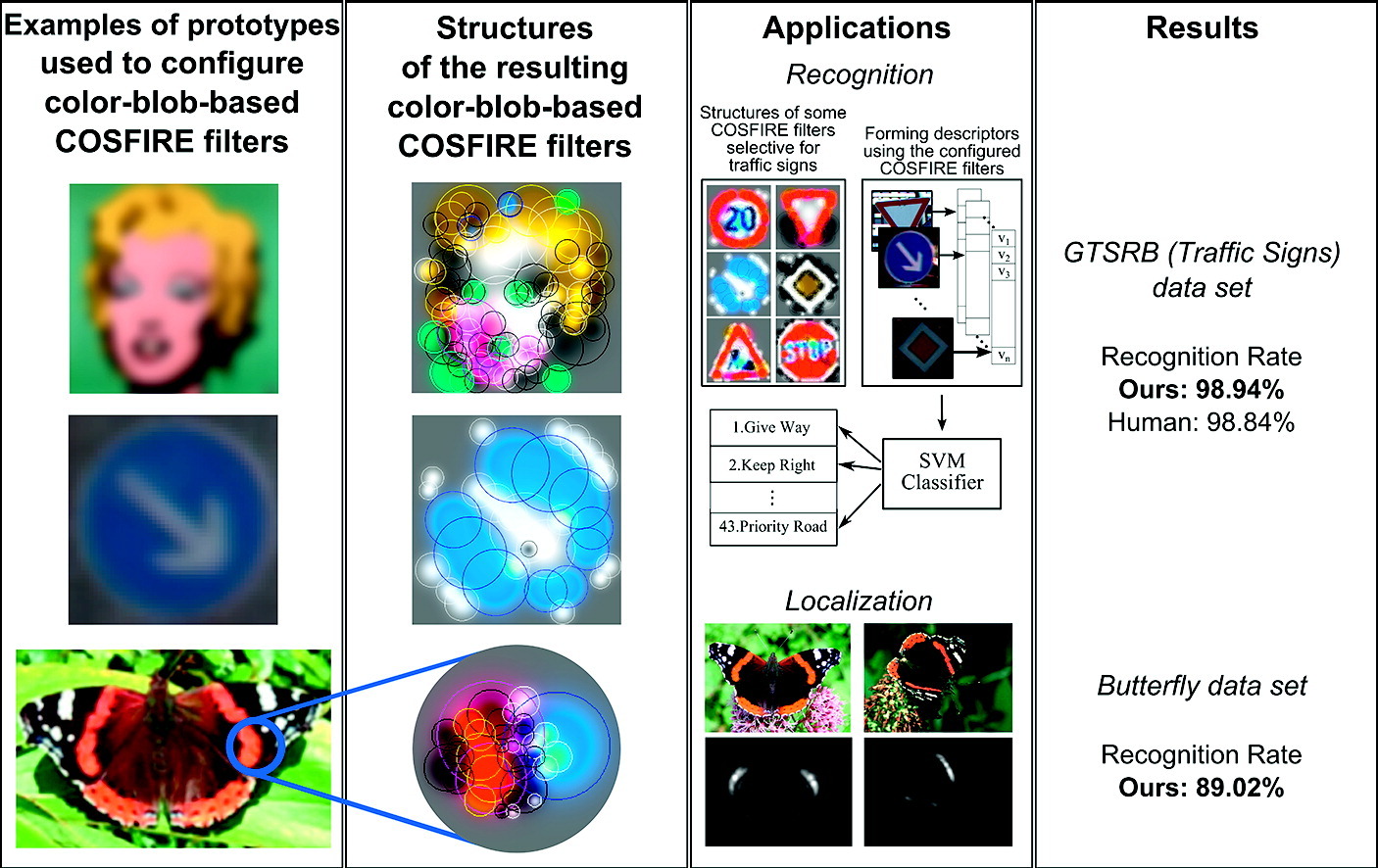
B. Gecer, G. Azzopardi, and N. Petkov, “Color-blob-based COSFIRE filters for Object Recognition” Image and Vision Computing, vol. 57, pp. 165-174, 2017.
Gecer, Baris. Detection and classification of breast cancer in whole slide histopathology images using deep convolutional networks. Diss. Bilkent University, 2016.